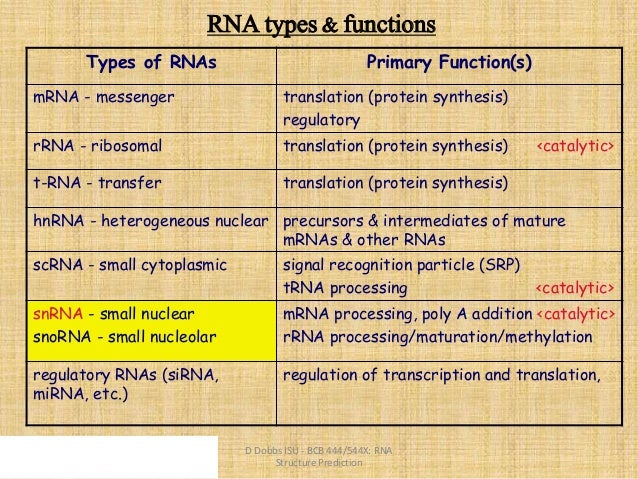
It has been shown via rigorous cross-validations for the RNA sequences from Homo sapiens and Mus musculus transcriptomes that the success rates achieved by the powerful new predictor are quite high. To address such a challenge, we developed a predictor called “iRNA-3typeA,” by which we can simultaneously identify the occurrence sites of the following three most frequently observed modifications in RNA: (1) N 1-methyladenosine (m 1A), (2) N 6-methyladenosine (m 6A), and (3) adenosine to inosine (A-to-I). However, so far no method whatsoever has been developed for simultaneously identifying several different types of RNA modifications. With the avalanche of RNA sequences generated in the post-genomic age, many computational methods have been proposed for identifying various types of RNA modifications one by one. Knowledge about the occurrence sites of these modifications is essential for in-depth understanding of the biological functions and mechanisms and for treating some genomic diseases as well. RNA modifications are additions of chemical groups to nucleotides or their local structural changes. Gene Editing: Technology & Applications. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
